Review Articles
M. Joannides1, A. N. Mays1, A. R. Mistry1, S. K. Hasan2, A. Reiter3, J. L. Wiemels4, C. A. Felix5, F. Lo Coco2, N. Osheroff6, E. Solomon1 and D. Grimwade1
2Department of Biopathology & Fondazione Santa Lucia, University of Tor Vergata, Rome, Italy.
3III.Medizinische Klinik,Universitätsmedizin Mannheim, Mannheim, Germany.
4Department of Epidemiology and Biostatistics, University of California, San Francisco, San Francisco, USA.
5Division of Oncology, Children’s Hospital of Philadelphia, Philadelphia, USA.
6The Departments of Biochemistry and Medicine, Vanderbilt University School of Medicine, Nashville, USA.
Published: October 24, 2011
Received: August 25, 2011
Accepted: September 20, 2011
Mediterr J Hematol Infect Dis 2011, 3(1): e2011045, DOI 10.4084/MJHID.2011.045
This article is available from: http://www.mjhid.org/article/view/9085
Abstract
Balanced chromosomal
translocations that generate chimeric oncoproteins
are considered to be initiating lesions in the pathogenesis of acute
myeloid leukemia. The most frequent is the t(15;17)(q22;q21), which
fuses the PML and RARA genes, giving rise to acute promyelocytic
leukemia (APL). An increasing proportion of APL cases are
therapy-related (t-APL), which develop following exposure to
radiotherapy and/or chemotherapeutic agents that target DNA
topoisomerase II (topoII), particularly mitoxantrone and epirubicin. To
gain insights into molecular mechanisms underlying the formation of the
t(15;17) we mapped the translocation breakpoints in a series of t-APLs,
which revealed significant clustering according to the nature of the
drug exposure. Remarkably, in approximately half of t-APL cases arising
following mitoxantrone treatment for breast cancer or multiple
sclerosis, the chromosome 15 breakpoint fell within an 8-bp “hotspot”
region in PML intron 6, which was confirmed to be a preferential site
of topoII-mediated DNA cleavage induced by mitoxantrone. Chromosome 15
breakpoints falling outside the “hotspot”, and the corresponding RARA
breakpoints were also shown to be functional topoII cleavage sites. The
observation that particular regions of the PML and RARA loci are
susceptible to topoII-mediated DNA damage induced by epirubicin and
mitoxantrone may underlie the propensity of these agents to cause APL.
Introduction
Acute
myeloid leukemia (AML) is characterized by a spectrum of recurring
chromosomal abnormalities, which distinguish biologically and
prognostically distinct subtypes of disease (reviewed [1]).
To date, more than one hundred balanced chromosomal rearrangements
(translocations, insertions and inversions) have been identified and
cloned,[2] with evidence suggesting that these are
critical initiating events in the pathogenesis of AML. Identification
of the genes involved in chromosomal rearrangements has provided major
insights into the regulation of normal hematopoiesis and how disruption
of key transcription factors and epigenetic modulators promote leukemic
transformation. A number of genes, including MLL at 11q23, NUP98 at
11p15, RARa at 17q21 and
RUNX1 at 21q22, are recurrent targets of chromosomal rearrangements in
AML, being fused to a range of potential partner genes (reviewed [3]).
Interestingly, chromosomal rearrangements involving these particular
loci also have been noted as a feature of secondary acute leukemias
arising as a complication of prior therapy involving drugs targeting
topoisomerase II (topoII), which are widely used in the treatment of a
variety of tumors.[4-13] TopoII is a critical enzyme
that relaxes supercoiled DNA and removes knots and tangles from the
genome by transiently cleaving and religating both strands of the
double helix via the formation of a covalent cleavage intermediate
(reviewed [14]). Epipodophyllotoxins (e.g.
etoposide), anthracyclines (e.g. epirubicin) and anthracenediones (e.g.
mitoxantrone) act as topoII poisons, inducing DNA damage by disrupting
the cleavage-religation equilibrium and increasing the concentration of
DNA topoII covalent complexes, which leads to apoptosis of the tumor
cells.[14]
The association between exposure to chemotherapeutic agents targeting
topoII and development of leukemias with balanced chromosomal
rearrangements has naturally implicated the enzyme in this process, but
the mechanisms involved have remained subject to debate. One hypothesis
takes into account reports that leukemia-associated translocations can
be detected in hematopoietic cells derived from healthy individuals
without overt leukemia,[15,16] suggesting that
administration of chemotherapy and/or radiotherapy provides a selective
advantage to progenitors with pre-existing translocations during
regrowth of depopulated bone marrow. Moreover, the exposure to DNA
damaging agents could lead to the acquisition of additional mutations
that cooperate with the chimeric fusion protein generated by the
translocation to induce leukemic transformation. A second hypothesis,
based on findings with transformed cells, raised the possibility that
agents targeting topoII could lead to the formation of chromosomal
translocations through an indirect mechanism involving induction of
apoptotic nucleases.[17-20] Interestingly, Rolf
Marschalek and colleagues provide evidence for a third potential
mechanism, showing that the region of the MLL locus where breakpoints
associated with infant and therapy-related leukemias cluster,
colocalize with an internal promoter element, highlighting the
relevance of chromatin structure and DNA topology in the genesis of
chromosomal translocations.[21] Finally, the fourth
hypothesis, which is based on increasing biochemical and genetic
evidence, suggests that in the presence of a topo II-targeting
chemotherapeutic agent, topoII plays a direct role in generating
double-stranded DNA breaks in regions of the genome that are
particularly susceptible due to the nature of the surrounding
chromatin. Following aberrant repair, these breaks go on to generate
leukemia-associated chromosomal translocations (reviewed [22]).
Intriguingly, the nature of the drug exposure has a bearing on the
molecular phenotype of the resultant secondary leukemia, with
translocations involving the MLL gene at 11q23 being particularly
associated with etoposide exposure.[10,23,24]
Development of therapy-related acute promyelocytic leukemia (t-APL),
characterized by the t(15;17)(q22;q21), has been linked to treatment
with mitoxantrone and epirubicin.[12,25,26]
The t(15;17) leads to fusion of the gene encoding the myeloid
transcription factor RARa
(Retinoic Acid Receptor Alpha) at 17q21 with a gene that was previously
unknown - designated PML (for ProMyelocytic Leukemia), which has
subsequently been found to be involved in growth suppression and
regulation of apoptosis (reviewed [27]). This subtype
of leukemia is of particular interest, being the first in which
molecularly targeted therapies (i.e., all-transretinoic acid [ATRA] and
arsenic trioxide [ATO]) have been successfully used in clinical
practice.[27] These agents act by inducing
degradation of the PML-RARa
oncoprotein, leading to clinical remission and have resulted in
dramatic improvements in clinical outcome (reviewed [28]).
They also offer a potentially curative approach in patients with t-APL
who have already received significant doses of chemotherapy for their
previous condition and may be close to the anthracycline ceiling, or
who are considered unfit for conventional therapy.[29]
The majority of t-APL cases arise in patients who have undergone
treatment for breast cancer, where mitoxantrone and epirubicin have
been widely used.[12,25,26,30]
In this setting, cumulative doses of epirubicin of ≤720mg/m2 have been
associated with a secondary leukemia risk of 0.37% at 8 years.[31] As more patients survive their primary cancers,
secondary leukemias are becoming an increasing healthcare problem.[32]
Although t-APL remains relatively uncommon, two case series have
suggested that the incidence has risen in recent years, with up to 20%
of APL patients presenting with secondary disease.[25,30]
Investigation of Molecular Mechanisms in Mitoxantrone-Related t-APL. As
a first step to gain insights into mechanisms underlying formation of
the t(15;17) we used long-range polymerase chain reaction (PCR) and
sequence analysis to define chromosomal breakpoint locations, comparing
the pattern between patients presenting de novo (n=35) and those with
t-APL occurring following exposure to mitoxantrone (n=6) or other
agents/radiation therapy (n=7).[33] Analysis of
diagnostic samples from large cohorts of patients with de novo APL has
established that the majority of chromosome 15 breakpoints fall within
3 breakpoint cluster regions (bcr) i.e. in intron 3 (bcr3), intron 6
(bcr1) and exon 6 (bcr2) of the PML locus, accounting for approximately
40%, 55% and 5% of cases respectively.[34] Chromosome
17 breakpoints fall within the ~17kb intron 2 of the RARa locus, such that the PML-RARa
fusion retains key functional domains mediating DNA binding and
interaction with coactivator/corepressors, retinoid-X-receptor and
ligand (i.e. retinoic acid).[27] While breakpoints
observed in de novo APL appeared broadly distributed, chromosome 15
breakpoints of each of the mitoxantrone-related t-APLs fell within PML
intron 6, with 4 cases clustering within an 8-bp region (position
1482-9)(see Figure 1).[33]
Given that this intron is over 1kb in length, this clustering of
breakpoints was highly unlikely to have occurred by chance (p<0.001
by scan statistics). To investigate this further, we used a functional
in vitro assay, in which substrates containing the normal homologues of
the PML and RARa
translocation breakpoints are 5’-end-labelled and exposed to clinically
relevant topoII poisons (e.g. mitoxantrone, epirubicin, etoposide) in
the absence or presence of human topoII alpha; cleavage complexes are
trapped and cleavage sites mapped in relation to the observed
translocation breakpoints at the sequence level.[35-37]
These experiments demonstrated that the breakpoint “hotspot,”
identified in t-APL patients previously treated with mitoxantrone for
breast cancer, corresponded precisely to a preferential
mitoxantrone-induced topoII-dependent DNA cleavage site at position
1484 (see Figure 2).[33] Moreover, each observed translocation breakpoint
within the RARa locus on
chromosome 17 was confirmed to be a preferred site of topoII-mediated
DNA damage induced by mitoxantrone (Figure
2).[33]
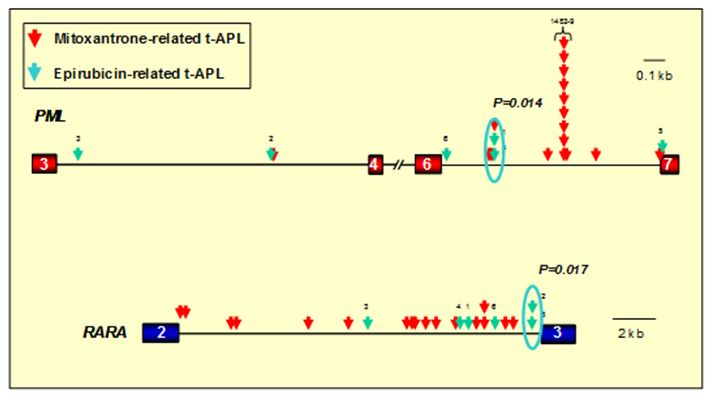
Figure 1. Distribution of translocation breakpoints within the PML and RARA loci in t-APL cases arising following epirubicin and mitoxantrone. PML exons are represented by red boxes, RARA exons in blue and introns are represented by black lines. Arrows indicate the location of PML and RARA translocation breakpoints identified by long-range PCR and sequence analysis in patients with t-APL arising following mitoxantrone (red arrows) or epirubicin (green arrows). In 12 patients mitoxantrone was used for treatment of multiple sclerosis (MS). In the remaining 5 patients with mitoxantrone-related APL and the 6 patients with t-APL following epirubicin, these agents were used for prior breast cancer. Significant breakpoint clustering was observed, with a “hotspot” identified in PML intron 6 (position 1482-9) in mitoxantrone-related APL (following use of the drug for MS or breast cancer) and separate clusters associated with APL arising following epirubicin exposure. Chromosomal breakpoints were confirmed to be preferential sites of drug-induced topoisomerase II cleavage in functional assays (see Figure 2). Adapted from Mays et al.[42] with permission.
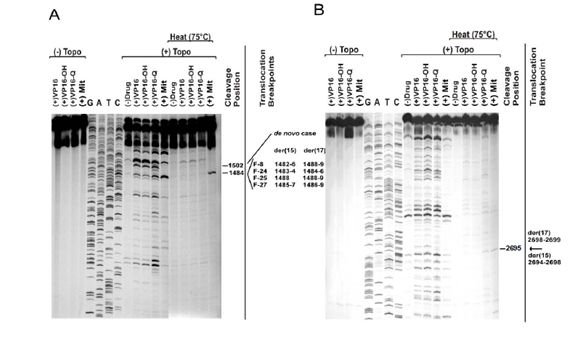
Figure 2. Demonstration of mitoxantrone-induced topoisomerase II dependent DNA cleavage at translocation breakpoints in therapy-related APL.
A) In vitro DNA topoisomerase II (topoII) cleavage assay carried out for a PML substrate containing the breakpoints of 4 treatment-related APL (t-APL) cases (F-8,-24,-25,-27) within the 8-bp breakpoint “hotspot” (positions 1482-1489). Patients received combination chemotherapy including the topoII poison mitoxantrone for breast cancer. Control reactions were carried out in the absence of topoII (lanes 1-4), and in the presence of etoposide (VP16), etoposide catechol (VP16-OH), etoposide quinone (VP16-Q) and mitoxantrone (Mit). Dideoxy sequencing reactions of the substrate are shown in lanes 5-8. Cleavage reactions were carried out by exposure to human topoIIa in the absence of drug (lane 9), and in the presence of etoposide (lane 10), etoposide catechol (lane 11), etoposide quinone (lane 12) and mitoxantrone (lane 13). Additional cleavage reactions were carried out to evaluate the heat-stability of cleavage complexes formed by incubation at 75°C for 10 min (lanes 14-18). The nucleotide shown by the dash is the 5’side of the cleavage site (-1 position), which corresponds to the der(15) and der(17) translocation breakpoints in 4 cases of mitoxantrone-related APL (far right). The cleavage site at position 1484 was observed in the absence of drug, and in the presence of etoposide, both etoposide metabolites and mitoxantrone (lanes 9-13). Cleavage at this position was the strongest site observed in the presence of mitoxantrone (lane 13). Furthermore, the complexes formed at this site were shown to be heat-stable in the presence of mitoxantrone (lane 18). Interestingly, a cleavage site at position 1502 is also observed, which corresponds to a breakpoint detected in a case of de novo APL.
B) TopoII cleavage assay of normal homologue of der(15) and der(17) RARA translocation breakpoints in APL of one of the mitoxantrone-related cases (F-8). The substrate spanning positions 2603 to 2871 of RARA intron 2 contained the translocation breakpoints. Dash at right shows (-1) position of cleavage site corresponding to der(15) and der(17) translocation breakpoints (arrow far right). Adapted from Mistry et al.[33] with permission.
These
data strongly implicated mitoxantrone in the etiology of t-APL.
However, it is important to consider that the study of patients
developing leukemia following cancer therapy presents a challenge,
given that they have often been exposed to multiple cytotoxic drugs in
addition to radiotherapy. This makes it difficult to identify the
causative agent with any certainty. Moreover, such patient populations
could feasibly be enriched for individuals at particular risk of
leukemia, having already developed one form of cancer. Therefore, to
provide further insights into molecular mechanisms in topoII-related
leukemias, we analyzed a cohort of 12 patients collected from across
Europe who developed APL following the use of single agent mitoxantrone
to treat a benign condition, multiple sclerosis (MS), and in whom there
was no history of previous cancer.[38] Chromosome 15
breakpoints again were found to cluster at position 1484 within PML
intron 6. Furthermore, shared chromosome 17 breakpoints that were
preferential sites of mitoxantrone-induced topoII cleavage in
functional assays were identified within RARAa intron 2 (Figure 1).[38]
The series of mitoxantrone-related t-APL cases analyzed has been
further extended recently, with the chromosome 15 breakpoint found to
fall within the “hotspot” region in 12 of 23 cases (52%).[33,38,39]
Comparison of the genomic breakpoint junction regions with the native
genes showed that translocations in mitoxantrone-related t-APL were
reciprocal, generally without loss or gain of nucleotides.[33,38] Presence of sequence homologies between PML and RARa
suggests that topoII-mediated DNA damage may be repaired by the
canonical nonhomologous end-joining (NHEJ) or the alternative
end-joining (alt-NHEJ) pathway, which require minimal overlapping
sequences between nonhomologous chromosomes to repair double- stranded
DNA breaks (reviewed [40]). Using the information
derived from genomic breakpoint junction sequence analysis and in vitro
cleavage assays, the knowledge that topoII introduces staggered nicks
in DNA with 4-bp overhangs 22 and considering known mechanisms of DNA
repair it was possible to construct models by which the t(15;17) may
have been formed in each case analyzed (see Figures 3 & 4). Taken together,
these data provide very strong evidence that mitoxantrone is a
causative agent in the pathogenesis of t-APL.
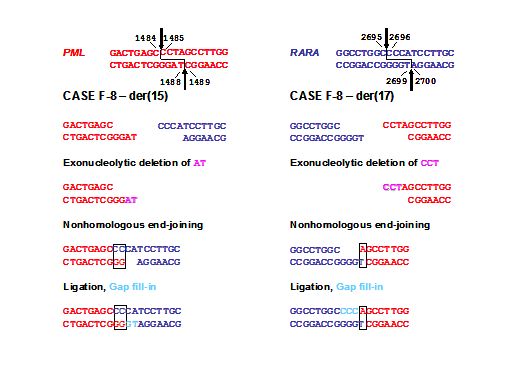
Figure 3. Model for formation of the t(15;17) in a case of mitoxantrone-related t-APL (case F8) following topoII induced cleavage in PML and RARA loci with 4-base 5’ overhangs and aberrant DNA repair. Native PML and RARA sequences are red and blue, respectively. The processing includes exonucleolytic nibbling to form two-base (der(15)) or single-base (der(17)) homologies and creation of both breakpoint junctions by error-prone nonhomologous end-joining (NHEJ). In formation of the der(15), positions 1487-1488 on the antisense strand of PML are lost by exonucleolytic nibbling (pink) before NHEJ joins the indicated bases. Positions 1485-1487 on the sense strand of PML are lost by exonucleolytic nibbling (pink) and the der(17) forms by NHEJ. Template-directed polymerization of the relevant single-stranded overhangs fills in any gaps (light blue). Each RARA overhang is preserved completely. Adapted from Mistry et al.[33] with permission.
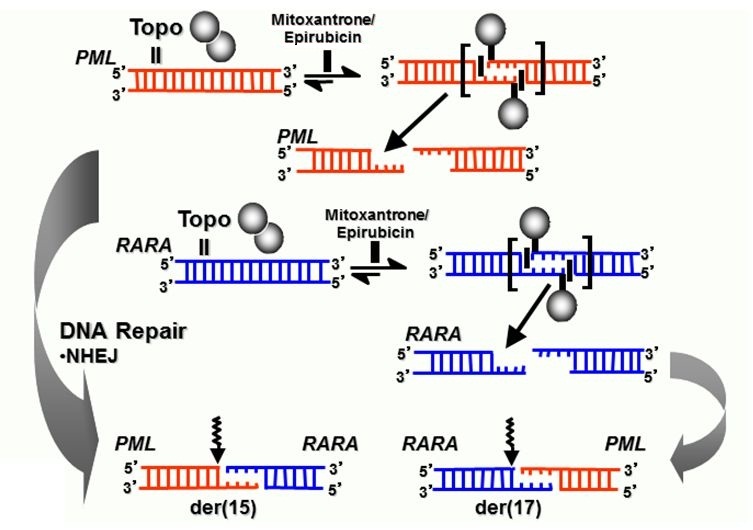
Figure 4. Model summarizing formation of reciprocal translocation breakpoint junctions in treatment related APL directly by generation of drug-stimulated topoisomerase II cleavage complexes and near-precise or precise NHEJ repair. Adapted from Felix et al.[22] with permission.
Investigation of Molecular Mechanisms in t-APL Cases Arising Following Other TopoII Poisons
Epirubicin exposure has been linked to secondary leukemias with a range of balanced rearrangements, including translocations involving the MLL locus, core binding factor leukemias and t-APL with the t(15;17).[31,41] In order to gain further insights into molecular mechanisms underlying epirubicin-related leukemias, we characterized t(15;17) genomic breakpoint junction regions in a series of 6 t-APL cases that arose following breast cancer therapy.[42] Epirubicin was generally used as a component of combination chemotherapy, with a median latency period from first exposure to presentation of t-APL of 26 months (range 18-48 months). Although the number of cases examined was small, significant breakpoint clustering was observed in both the PML and RARA loci (P= .009 and P = .017, respectively), with PML breakpoints lying outside the mitoxantrone-associated “hotspot” region (Figure 1). Functional assays demonstrated that recurrent breakpoints identified in the PML and RARA loci in epirubicin-related t-APL were preferential sites of topoII-induced DNA damage that were enhanced by epirubicin.[42] Again, using the same approach as for mitoxantrone-related t-APLs, models could be constructed to explain the formation of the t(15;17) in APLs arising following epirubicin exposure.[42] There also have been reports of t-APL occurring following treatment with other topoII poisons (e.g., etoposide) used for lymphomas and various solid tumors, as well as Langerhans cell histiocytosis.[12,25,43] To determine whether topoII–mediated cleavage is relevant to other drugs associated with t-APL, we also have studied a patient in whom APL developed after treatment for laryngeal carcinoma that included etoposide and doxorubicin.[33] Etoposide and its metabolites and doxorubicin induced topoII to cleave DNA at the PML and RARa translocation breakpoints. Moreover, the cleavage sites could recombine to form the der(15) and der(17) breakpoint junctions observed in this patient. Taken together, these results suggest that topoII–mediated cleavage is a general mechanism causing DNA damage in APL that develops after treatment with various agents that target topoII (Figure 4).
Concluding Remarks
While therapy-related leukemias are still relatively uncommon, they are
characterized by the same range of cytogenetic abnormalities that are
found in cases of AML arising de novo.[44,45] Indeed,
greater understanding of therapy-related leukemias may provide
significant insights into the biology of their de novo counterparts.
For example, defining the latency period between first exposure to a
leukemogenic agent (e.g. mitoxantrone) and presentation with full blown
leukemia, provides clues to the timeframe between acquisition of
chromosomal rearrangements such as the t(15;17) and progression to
leukemia in the de novo setting. Analysis of t-APL cases suggests that
the median time to develop APL from the formation of the t(15;17) is
~27 months,[33,38,42]
implying the need for cooperating mutations. While logistically
challenging, therapy-related leukemias afford the opportunity for
tracking the stepwise acquisition of mutations that are required for
progression to full-blown leukemia,[46,47] and which
may be of relevance to the pathogenesis of leukemias arising de novo.
While we have observed a few cases of t-APL that present within 12
months from first mitoxantrone exposure, latency periods in the
majority of cases are much longer. This may account for why, even if
the t(15;17) were acquired in some cases in utero, de novo pediatric
APL only very rarely presents in infancy.
A number of studies conducted over the last three decades have served
to identify specific dosing schedules or particular agents that are
associated with high rates of induction of secondary leukemias,[32,48-51]] leading to the development
of effective alternative treatment protocols that are substantially
safer.[52-54]
However, the study of patients with t-APL has demonstrated that
therapy-related leukemias also can occur in patients subject to very
low level exposure, as exemplified by a case of APL involving the PML
breakpoint “hotspot” arising following a single 15 mg dose of
mitoxantrone used as adjuvant chemotherapy for breast cancer.[33]
Epidemiology studies conducted in MS patients treated with mitoxantrone
have suggested that the risk of development of secondary leukemia is ~1
in 370, [55,56] with the majority of reported cases
being t-APL. This raises key questions as to the extent to which the
play of chance is involved in the development of therapy-related
leukemias, as well as the relative importance of patient predisposition
to the development of this complication. A number of prerequisites have
to be satisfied to develop leukemia following treatment with a topoII
poison. Firstly, double-strand DNA breaks must occur within two genes
with the potential to form oncogenic fusions. These breaks then need to
be repaired to generate in-frame functional chimeric fusion genes. This
translocation event needs to occur in a progenitor permissive for
leukemic transformation and finally the necessary cooperating mutations
are acquired. While our studies have provided evidence that topoII
plays a direct role in mediating DNA damage that leads to formation of
the t(15;17) in t-APL, a key question remains as to whether the enzyme
is also involved in the formation of chromosomal translocations in de
novo leukemias. Exposure to environmental toxins and agents targeting
topoII has been implicated in the development of infant leukemia with
translocations involving MLL at 11q23.[57-59]
Interestingly, recent evidence lends further support for topoII in the
etiology of chromosomal translocations, inducing DNA damage in the
TMPRSS2 and ERG loci in response to androgen signalling, leading to
formation of fusion genes involved in prostate cancer.[60]
It is readily conceivable that genetic susceptibility to primary tumors
due to mutations or functional variants for example in DNA repair
pathways also could increase the risk of development of secondary
leukemias. Interestingly, a recent genome-wide association study has
implicated variants in the PRDM1 gene at 6q21 in the development of
second neoplasms in children treated with radiotherapy for Hodgkin’s
disease,[61] whereas whole genome sequencing applied
in a case of therapy-related AML arising from early-onset BRCA1/2
mutation-negative breast and ovarian cancer revealed a novel TP53
cancer susceptibility mutation.[62] The spectrum of
resultant leukemias could reflect the nature of the genetic
susceptibility as well as the agents preferentially used for the
particular primary condition, as would be suggested by the propensity
of etoposide to induce secondary leukemias involving the MLL gene at
11q23 and epirubicin and mitoxantrone to induce t-APL. Moreover,
genetic variation in the handling of a range of specific cytotoxic
agents could affect an individual’s risk of developing secondary
leukemia (reviewed [63]). Indeed, it has recently
been reported that genetic variants of genes encoding drug-metabolizing
enzymes and components of DNA repair pathways are associated with
increased susceptibility to development of t-APL in patients with MS
receiving mitoxantrone.[64]
Dissecting out the relative importance of these factors represents a
considerable challenge. It requires the analysis of substantial patient
cohorts, which are well characterized both in terms of their primary
tumors, prior cytotoxic therapy and cytogenetic and molecular profile
of the secondary leukemias. Nevertheless, significant progress in this
research area is likely to be fruitful allowing not only the
development of more individualized and safer approaches to treatment of
primary tumors, but also (potentially) providing insights into
molecular mechanisms underlying the pathogenesis of de novo leukemias.
Thus, it could afford improved understanding of AML as a whole.
Acknowledgements
DG gratefully acknowledges Leukaemia & Lymphoma
Research for support, with award of Gordon Piller Studentships to
Ashley Mays and Melanie Joannides. We also acknowledge support from the
Genetics theme of the Guy’s and St. Thomas’ NHS Foundation Trust
National Institute for Health Research (NIHR) Biomedical Research
Centre. NO acknowledges support by National Institutes of Health grant
GM033944.
References
- Smith ML, Hills RK, Grimwade D. Independent
prognostic variables in acute myeloid leukaemia. Blood Rev 2011;
25:39-51. http://dx.doi.org/10.1016/j.blre.2010.10.002
PMid:21078537

- Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer (2011). Mitelman F, Johansson B, and Mertens F. (Eds.), http://cgap.nci.nih.gov/Chromosomes/Mitelman
- Zhang Y, Rowley JD. Chromatin structural
elements and chromosomal translocations in leukemia. DNA Repair (Amst).
2006; 5:1282-97. http://dx.doi.org/10.1016/j.dnarep.2006.05.020
PMid:16893685

- Ahuja HG, Felix CA, Aplan PD. Potential
role of DNA topoisomerase ll poisons in the generation of
t(11;20)(p15;q11) translocations. Genes Chromosomes Cancer 2000;
29:96-105. http://dx.doi.org/10.1002/1098-2264(2000)9999:9999<::AID-GCC1013>3.0.CO;2-T

- Pedersen-Bjergaard J, Andersen, MK,
Christiansen DH, Nerlov C. Genetic pathways in therapy-related
myelodysplasia and acute myeloid leukemia. Blood 2002; 99:1909-1912. http://dx.doi.org/10.1182/blood.V99.6.1909
PMid:11877259

- Pedersen-Bjergaard, J, Andersen, MK,
Andersen, MT, Christiansen DH. Genetics of therapy related
myelodysplasia and acute myeloid leukemia. Leukemia 2008; 22: 240-248. http://dx.doi.org/10.1038/sj.leu.2405078
PMid:18200041

- Rowley JD, Olney H. International workshop
on the relationship of prior therapy to balanced chromosome aberrations
in therapy related myelodysplastic syndromes and acute leukemia:
overview report. Genes Chromosomes Cancer 2002; 33:331-345. http://dx.doi.org/10.1002/gcc.10040
PMid:11921269

- Larson RA, Le Beau MM. Therapy-related
myeloid leukaemia: a model for leukemogenesis in humans. Chem Biol
Interact. 2005; 153-154:187-95. http://dx.doi.org/10.1016/j.cbi.2005.03.023
PMid:15935816

- Allan JM, Travis LB. Mechanisms of therapy
related carcinogenesis. Nature Reviews 2005; 5:943-955. http://dx.doi.org/10.1038/nrc1749
PMid:16294218

- Bloomfield CD, Archer KJ, Mrózek K,
Lillington DM, Kaneko Y, Head DR, Dal Cin P, Raimondi SC. 11q23
balanced chromosome aberrations in treatment-related myelodysplastic
syndromes and acute leukemia: report from an International Workshop.
Genes Chromosomes Cancer 2002; 33:362-78. http://dx.doi.org/10.1002/gcc.10046
PMid:11921271

- Slovak ML, Bedell V, Popplewell L, Arber
DA, Schoch C, Slater R. 21q22 balanced chromosome aberrations in
therapy-related hematopoietic disorders: report from an International
Workshop. Genes Chromosomes Cancer 2002; 33:379-94. http://dx.doi.org/10.1002/gcc.10042
PMid:11921272

- Andersen MK, Larson RA, Mauritzson N,
Schnittger S, Jhanwar SC, Pedersen-Bjergaard J. Balanced chromosome
abnormalities inv(16) and t(15;17) in therapy-related myelodysplastic
syndromes and acute leukemia: report from an international workshop.
Genes Chromosomes Cancer. 2002; 33:395-400. http://dx.doi.org/10.1002/gcc.10043
PMid:11921273

- Romana SP, Radford-Weiss I, Ben Abdelali
R, Schluth C, Petit A, Dastugue N, Talmant P, Bilhou-Nabera C, Mugneret
F, Lafage-Pochitaloff M, Mozziconacci MJ, Andrieu J, Lai JL, Terre C,
Rack K, Cornillet-Lefebvre P, Luquet I, Nadal N, Nguyen-Khac F, Perot
C, Van den Akker J, Fert-Ferrer S, Cabrol C, Charrin C, Tigaud I,
Poirel H, Vekemans M, Bernard OA, Berger R; Groupe Francophone de
Cytogénétique Hématologique. NUP98 rearrangements in hematopoietic
malignancies: a study of the Groupe Francophone de Cytogénétique
Hématologique. Leukemia. 2006; 20:696-706. http://dx.doi.org/10.1038/sj.leu.2404130
PMid:16467868

- Deweese JE, Osheroff N. The DNA cleavage
reaction of topoisomerase II: wolf in sheep's clothing. Nucleic Acids
Res. 2009; 37:738-48. http://dx.doi.org/10.1093/nar/gkn937 PMid:19042970
PMCid:2647315

- Mori H, Colman SM, Xiao Z, Ford AM, Healy
LE, Donaldson C, Hows JM, Navarrete C, Greaves M. Chromosome
translocations and covert leukemic clones are generated during normal
fetal development. Proc Natl Acad Sci U S A 2002; 99: 8242-8247. http://dx.doi.org/10.1073/pnas.112218799
PMid:12048236 PMCid:123052

- Basecke J, Cepek L, Mannhalter C, Krauter
J, Hildenhagen S, Brittinger G, Trumper L, Griesinger F. Transcription
of AML1/ETO in bone marrow and cord blood of individuals without acute
myelogenous leukemia. Blood 2002; 100:2267-2268. http://dx.doi.org/10.1182/blood-2002-06-1673
PMid:12229881

- Stanulla M, Wang J, Chervinsky DS, Thandla
S, Aplan PD. DNA cleavage within the MLL breakpoint cluster region is a
specific event which occurs as part of higher-order chromatin
fragmentation during the initial stages of apoptosis. Mol Cell Biol
1997; 17:4070-4079. PMid:9199342 PMCid:232260

- Betti CJ, Villalobos MJ, Diaz MO, Vaughan
AT. Apoptotic triggers initiate translocations within the MLL gene
involving nonhomologous end joining repair system. Cancer Res 2001;
61:4550-4555. PMid:11389089

- Betti CJ, Villalobos MJ, Diaz MO, Vaughan
AT. Apoptotic stimuli initiate MLL-AF9 translocations that are
transcribed in cells capable of division. Cancer Res 2003; 63:
1377-1381. PMid:12649202

- Sim SP, Liu LF. Nucleolytic cleavage of
the mixed lineage leukemia breakpoint cluster region during apoptosis.
J Biol Chem 2001; 276:31590-31595. http://dx.doi.org/10.1074/jbc.M103962200
PMid:11406628

- Scharf S, Zech J, Bursen A, Schraets D,
Oliver PL, Kliem S, Pfitzner E, Gillert E, Dingermann T, Marschalek R.
Transcription linked to recombination: a gene-internal promoter
coincides with the recombination hot spot II of the human MLL gene.
Oncogene. 2007; 26:1361-71. http://dx.doi.org/10.1038/sj.onc.1209948
PMid:16983345

- Felix CA, Kolaris CP, Osheroff N.
Topoisomerase II and the etiology of chromosomal translocations. DNA
Repair (Amst) 2006; 5:1093-108. http://dx.doi.org/10.1016/j.dnarep.2006.05.031
PMid:16857431

- Andersen MK, Johansson B, Larsen SO,
Pedersen-Bjergaard J. Chromosomal abnormalities in secondary MDS and
AML. Relationship to drugs and radiation with specific emphasis on the
balanced rearrangements. Haematologica. 1998; 83:483-8. PMid:9676019

- Sung PA, Libura J, Richardson C. Etoposide
and illegitimate DNA double-strand break repair in the generation of
MLL translocations: new insights and new questions. DNA Repair (Amst).
2006; 5:1109-18. http://dx.doi.org/10.1016/j.dnarep.2006.05.018
PMid:16809075

- Beaumont M, Sanz M, Carli PM, Maloisel F,
Thomas X, Detourmignies L, Guerci A, Gratecos N, Rayon C, San Miguel J,
Odriozola J, Cahn JY, Huguet F, Vekhof A, Stamatoulas A, Dombret H,
Capote F, Esteve J, Stoppa AM, Fenaux P. Therapy-related acute
promyelocytic leukemia. J Clin Oncol 2003; 21:2123-37. http://dx.doi.org/10.1200/JCO.2003.09.072
PMid:12775738

- Pulsoni A, Pagano L, Lo Coco F, Avvisati
G, Mele L, Di Bona E, Invernizzi R, Leoni F, Marmont F, Mele A, Melillo
L, Nosari AM, Pogliani EM, Vignetti M, Visani G, Zagonel V, Leone G,
Mandelli F. Clinicobiological features and outcome of acute
promyelocytic leukemia occurring as a second tumor: the GIMEMA
experience. Blood. 2002;100:1972-6. http://dx.doi.org/10.1182/blood-2001-12-0312
PMid:12200354

- Mistry AR, Pedersen EW, Solomon E,
Grimwade D. The molecular pathogenesis of acute promyelocytic
leukaemia: implications for the clinical management of the disease.
Blood Rev. 2003; 17:71-97. http://dx.doi.org/10.1016/S0268-960X(02)00075-9

- de Thé H, Chen Z. Acute promyelocytic
leukaemia: novel insights into the mechanisms of cure. Nat Rev Cancer.
2010; 10:775-83. http://dx.doi.org/10.1038/nrc2943 PMid:20966922

- Sanz MA, Grimwade D, Tallman MS, Lowenberg
B, Fenaux P, Estey EH, Naoe T, Lengfelder E, Büchner T, Döhner H,
Burnett AK, Lo-Coco F. Management of acute promyelocytic leukemia:
Recommendations from an expert panel on behalf of the European
LeukemiaNet. Blood 2009; 113:1875-1891. http://dx.doi.org/10.1182/blood-2008-04-150250
PMid:18812465

- Carli PM, Sgro C, Parchin-Geneste N,
Isambert N, Mugneret F, Girodon F, Maynadie M. Increase therapy-related
leukemia secondary to breast cancer. Leukemia 2000; 14:1014-7. http://dx.doi.org/10.1038/sj.leu.2401787
PMid:10865966

- Praga C, Bergh J, Bliss J, Bonneterre J,
Cesana B, Coombes RC, Fargeot P, Folin A, Fumoleau P, Giuliani R,
Kerbrat P, Hery M, Nilsson J, Onida F, Piccart M, Shepherd L, Therasse
P, Wils J, Rogers D.Risk of acute myeloid leukemia and myelodysplastic
syndrome in trials of adjuvant epirubicin for early breast cancer:
Correlation with doses of epirubicin and cyclophosphamide. J Clin Oncol
2005; 23: 4179-4191. http://dx.doi.org/10.1200/JCO.2005.05.029
PMid:15961765

- Leone G, Luca M, Alessandro P, Equitani F,
Pagano, L. The incidence of secondary leukaemias. Hematologica 1999;
84:937-945.

- Mistry AR, Felix CA, Whitmarsh RJ, Mason
A, Reiter A, Cassinat B, Parry A, Walz C, Wiemels JL, Segal MR, Adès L,
Blair IA, Osheroff N, Peniket AJ, Lafage-Pochitaloff M, Cross NC,
Chomienne C, Solomon E, Fenaux P, Grimwade D. DNA topoisomerase II in
therapy-related acute promyelocytic leukemia. N Engl J Med 2005;
352:1529-38. http://dx.doi.org/10.1056/NEJMoa042715
PMid:15829534

- Grimwade D, Jovanovic JV, Hills RK, Nugent
EA, Patel Y, Flora R, Diverio D, Jones K, Aslett H, Batson E, Rennie K,
Angell R, Clark RE, Solomon E, Lo-Coco F, Wheatley K, Burnett AK.
Prospective minimal residual disease monitoring to predict relapse of
acute promyelocytic leukemia and to direct pre-emptive arsenic trioxide
therapy. J Clin Oncol, 2009; 27:3650-8. http://dx.doi.org/10.1200/JCO.2008.20.1533
PMid:19506161

- Lovett BD, Strumberg D, Blair IA, Pang S,
Burden DA, Megonigal MD, Rappaport EF, Rebbeck TR, Osheroff N, Pommier
YG, Felix CA. Etoposide metabolites enhance DNA topoisomerase II
cleavage near leukemia-associated MLL translocation breakpoints.
Biochemistry 2001; 40:1159-1170. http://dx.doi.org/10.1021/bi002361x
PMid:11170441

- Lovett BD, Lo Nigro L, Rappaport EF, Blair
IA, Osheroff N, Zheng N, Megonigal MD, Williams WR, Nowell PC, Felix
CA. Near-precise interchromosomal recombination and functional DNA
topoisomerase II cleavage sites at MLL and AF-4 genomic breakpoints in
treatment-related acute lymphoblastic leukemia with t(4;11)
translocation. Proc Natl Acad Sci USA 2001; 98:9802-9807. http://dx.doi.org/10.1073/pnas.171309898
PMid:11493704 PMCid:55533

- Whitmarsh RJ, Saginario C, Zhuo Y,
Hilgenfeld E, Rappaport EF, Megonigal MD, Carroll M, Liu M, Osheroff N,
Cheung NK, Slater DJ, Ried T, Knutsen T, Blair IA, Felix CA. Reciprocal
DNA topoisomerase II cleavage events at 5’-TATTA-3’ sequences in MLL
and AF-9 create homologous single-stranded overhangs that anneal to
form der(11) and der(9) genomic breakpoint junctions in
treatment-related AML without further processing. Oncogene 2003;
22:8448-59. http://dx.doi.org/10.1038/sj.onc.1207052
PMid:14627986

- Hasan SK, Mays AN, Ottone T, Ledda A, La
Nasa G, Cattaneo C, Borlenghi E, Melillo L, Montefusco E, Cervera J,
Stephen C, Satchi G, Lennard A, Libura M, Byl JA, Osheroff N, Amadori
S, Felix CA, Voso MT, Sperr WR, Esteve J, Sanz MA, Grimwade D, Lo Coco
F. Molecular analysis of t(15;17) genomic breakpoints in secondary
acute promyelocytic leukemia arising after treatment of multiple
sclerosis. Blood 2008; 112:3383-3390. http://dx.doi.org/10.1182/blood-2007-10-115600
PMid:18650449 PMCid:2954750

- Hasan SK, Ottone T, Schlenk RF, Xiao Y,
Wiemels JL, Mitra ME, Bernasconi P, Di Raimondo F, Stanghellini MT,
Marco P, Mays AN, Döhner H, Sanz MA, Amadori S, Grimwade D, Lo-Coco F.
Analysis of t(15;17) chromosomal breakpoint sequences in
therapy-related versus de novo acute promyelocytic leukemia:
association of DNA breaks with specific DNA motifs at PML and RARA
loci. Genes Chromosomes Cancer. 2010; 49:726-32. http://dx.doi.org/10.1002/gcc.20783
PMid:20544846

- Kass EM, Jasin M. Collaboration and
competition between DNA double-strand break repair pathways. FEBS Lett.
2010; 584:3703-8. http://dx.doi.org/10.1016/j.febslet.2010.07.057
PMid:20691183

- Pedersen-Bjergaard J, Sigsgaard TC,
Nielsen D, Gjedde SB, Philip P, Hansen M, Larsen SO, Rørth M, Mouridsen
H, Dombernowsky P. Acute monocytic or myelomonocytic leukemia with
balanced chromosome translocations to band 11q23 after therapy with
4-epi-doxorubicin and cisplatin or cyclophosphamide for breast cancer.
J Clin Oncol. 1992; 10:1444-51. PMid:1517787

- Mays AN, Osheroff N, Xiao Y, Wiemels JL,
Felix CA, Byl JAW, Saravanamuttu K, Peniket A, Corser R, Chang C, Hoyle
C, Parker AN, Hasan SK, Lo-Coco F, Solomon E, Grimwade D. Evidence for
direct involvement of epirubicin in the formation of chromosomal
translocations in t(15;17) therapy-related acute promyelocytic
leukemia. Blood 2010; 115:326-30. http://dx.doi.org/10.1182/blood-2009-07-235051
PMid:19884644 PMCid:2808156

- Kudo K, Yoshida H, Kiyoi H, Numata S,
Horibe K, Naoe T. Etoposide-related acute promyelocytic leukemia.
Leukemia. 1998; 12:1171-5. http://dx.doi.org/10.1038/sj.leu.2401089
PMid:9697869

- Smith SM, Le Beau MM, Huo D, Karrison T,
Sobecks RM, Anastasi J, Vardiman JW, Rowley JD, Larson RA.
Clinical-cytogenetic associations in 306 patients with therapy-related
myelodysplasia and myeloid leukemia: the University of Chicago series.
Blood. 2003; 102:43-52. http://dx.doi.org/10.1182/blood-2002-11-3343
PMid:12623843

- Kayser S, Döhner K, Krauter J, Köhne CH,
Horst HA, Held G, von Lilienfeld-Toal M, Wilhelm S, Kündgen A, Götze K,
Rummel M, Nachbaur D, Schlegelberger B, Göhring G, Späth D, Morlok C,
Zucknick M, Ganser A, Döhner H, Schlenk RF; German-Austrian AMLSG. The
impact of therapy-related acute myeloid leukemia (AML) on outcome in
2853 adult patients with newly diagnosed AML. Blood. 2011; 117:2137-45.
http://dx.doi.org/10.1182/blood-2010-08-301713
PMid:21127174

- Megonigal MD, Cheung NK, Rappaport EF,
Nowell PC, Wilson RB, Jones DH, Addya K, Leonard DG, Kushner BH,
Williams TM, Lange BJ, Felix CA. Detection of leukemia-associated
MLL-GAS7 translocation early during chemotherapy with DNA topoisomerase
II inhibitors. Proc Natl Acad Sci U S A. 2000; 97:2814-9. http://dx.doi.org/10.1073/pnas.050397097
PMid:10706619

- Robinson BW, Cheung NK, Kolaris CP,
Jhanwar SC, Choi JK, Osheroff N, Felix CA. Prospective tracing of
MLL-FRYL clone with low MEIS1 expression from emergence during
neuroblastoma treatment to diagnosis of myelodysplastic syndrome.
Blood. 2008; 111:3802-12. http://dx.doi.org/10.1182/blood-2007-07-096065
PMid:18195096 PMCid:2275033

- Pedersen-Bjergaard J. Long-term
complications of cancer chemotherapy. J Clin Oncol. 1995; 13:1534-6.
PMid:7602341

- Smith MA, Rubinstein L, Ungerleider RS.
Therapy-related acute myeloid leukemia following treatment with
epipodophyllotoxins: estimating the risks. Med Pediatr Oncol. 1994;
23:86-98. http://dx.doi.org/10.1002/mpo.2950230205 PMid:8202047

- Smith RE. Risk for the development of
treatment-related acute myelocytic leukemia and myelodysplastic
syndrome among patients with breast cancer: review of the literature
and the National Surgical Adjuvant Breast and Bowel Project experience.
Clin Breast Cancer. 2003; 4:273-9. http://dx.doi.org/10.3816/CBC.2003.n.032
PMid:14651772

- Leone G, Pagano L, Ben-Yehuda D, Voso MT.
Therapy-related leukemia and myelodysplasia: susceptibility and
incidence. Haematologica. 2007; 92:1389-98. http://dx.doi.org/10.3324/haematol.11034
PMid:17768113

- Pui CH, Pei D, Sandlund JT, Ribeiro RC,
Rubnitz JE, Raimondi SC, Onciu M, Campana D, Kun LE, Jeha S, Cheng C,
Howard SC, Metzger ML, Bhojwani D, Downing JR, Evans WE, Relling MV.
Long-term results of St Jude Total Therapy Studies 11, 12, 13A, 13B,
and 14 for childhood acute lymphoblastic leukemia. Leukemia. 2010;
24:371-82. http://dx.doi.org/10.1038/leu.2009.252
PMid:20010620 PMCid:2820159

- Kushner BH, Kramer K, Modak S, Qin LX,
Yataghena K, Jhanwar SC, Cheung NK. Reduced risk of secondary leukemia
with fewer cycles of dose-intensive induction chemotherapy in patients
with neuroblastoma. Pediatr Blood Cancer. 2009; 53:17-22. http://dx.doi.org/10.1002/pbc.21931

- Felix CA. A safer regimen for high-risk
neuroblastoma. Pediatr Blood Cancer. 2009; 53:3-6. http://dx.doi.org/10.1002/pbc.22020

- Ghalie RG, Mauch E, Edan G, Hartung HP,
Gonsette RE, Eisenmann S, Le Page E, Butine MD, De Goodkin DE. A study
of therapy-related acute leukaemia after mitoxantrone therapy for
multiple sclerosis. Mult Scler 2002; 8:441-445. http://dx.doi.org/10.1191/1352458502ms836oa
PMid:12356214

- Ellis R, Boggild M. Therapy-related acute
leukaemia with Mitoxantrone: what is the risk and can we minimise it?
Mult Scler. 2009; 15:505-8. http://dx.doi.org/10.1177/1352458508100967
PMid:19251838

- Alexander FE, Patheal SL, Biondi A,
Brandalise S, Cabrera ME, Chan LC, Chen Z, Cimino G, Cordoba JC, Gu LJ,
Hussein H, Ishii E, Kamel AM, Labra S, Magalhães IQ, Mizutani S,
Petridou E, de Oliveira MP, Yuen P, Wiemels JL, Greaves MF.
Transplacental chemical exposure and risk of infant leukemia with MLL
gene fusion. Cancer Res 2001; 61:2542-6. PMid:11289128

- Hall GW. Childhood myeloid leukaemias.
Best Pract Res Clin Haematol 2001; 14:573-91. http://dx.doi.org/10.1053/beha.2001.0155

- Spector LG, Xie Y, Robison LL, Heerema NA,
Hilden JM, Lange B, Felix CA, Davies SM, Slavin J, Potter JD, Blair CK,
Reaman GH, Ross JA. Maternal diet and infant leukemia: the DNA
topoisomerase II inhibitor hypothesis: a report from the children's
oncology group. Cancer Epidemiol Biomarkers Prev. 2005; 14:651-5. http://dx.doi.org/10.1158/1055-9965.EPI-04-0602

- Haffner MC, Aryee MJ, Toubaji A, Esopi DM,
Albadine R, Gurel B, Isaacs WB, Bova GS, Liu W, Xu J, Meeker AK, Netto
G, De Marzo AM, Nelson WG, Yegnasubramanian S. Androgen-induced
TOP2B-mediated double-strand breaks and prostate cancer gene
rearrangements. Nat Genet 2010; 42:668-75. http://dx.doi.org/10.1038/ng.613
PMid:20601956 PMCid:3157086

- Best T, Li D, Skol AD, Kirchhoff T,
Jackson SA, Yasui Y, Bhatia S, Strong LC, Domchek SM, Nathanson KL,
Olopade OI, Huang RS, Mack TM, Conti DV, Offit K, Cozen W, Robison LL,
Onel K. Variants at 6q21 implicate PRDM1 in the etiology of
therapy-induced second malignancies after Hodgkin's lymphoma. Nat Med
2011; 17:941-3. http://dx.doi.org/10.1038/nm.2407 PMid:21785431

- Link DC, Schuettpelz LG, Shen D, Wang J,
Walter MJ, Kulkarni S, Payton JE, Ivanovich J, Goodfellow PJ, Le Beau
M, Koboldt DC, Dooling DJ, Fulton RS, Bender RH, Fulton LL, Delehaunty
KD, Fronick CC, Appelbaum EL, Schmidt H, Abbott R, O'Laughlin M, Chen
K, McLellan MD, Varghese N, Nagarajan R, Heath S, Graubert TA, Ding L,
Ley TJ, Zambetti GP, Wilson RK, Mardis ER. Identification of a novel
TP53 cancer susceptibility mutation through whole-genome sequencing of
a patient with therapy-related AML. JAMA 2011; 305:1568-76. http://dx.doi.org/10.1001/jama.2011.473
PMid:21505135 PMCid:3170052

- Seedhouse C, Russell N. Advances in the
understanding of susceptibility to treatment-related acute myeloid
leukaemia. Br J Haematol 2007; 137: 513-529. http://dx.doi.org/10.1111/j.1365-2141.2007.06613.x
PMid:17539774

- Hasan SK, Buttari F, Ottone T, Voso MT,
Hohaus S, Marasco E, Mantovani V, Garagnani P, Sanz MA, Cicconi L,
Bernardi G, Centonze D, Lo-Coco F. Risk of acute promyelocytic leukemia
in multiple sclerosis: coding variants of DNA repair genes. Neurology.
2011; 76:1059-65 http://dx.doi.org/10.1212/WNL.0b013e318211c3c8
PMid:21346221
